SELECTED PROJECTS:
PHYLOGENETICS AND EVOLUTION OF PARASITIC WASPS AND OTHER HYMENOPTERA
Ichneumonoidea is an exceptionally diverse (>3% of all life on the planet) group of parasitoid wasps that have successfully colonized numerous host lineages at different life stages and evolved several mechanisms to overcome host immune systems. I am currently examining how parasitic strategies in Ichneumonoidea (including host association, host switching, host stage attacked, and viral symbiosis) evolve over macroevolutionary timescales and how this relates to morphology, distribution, and speciation.

I am interested in cutting-edge genomic approaches to reconstruct robust phylogenies that allow for testing of macroevolutionary questions. See our recent papers here on evolution of Braconidae, Ichneumonoidea, Vespidae, and Evaniidae.
MOBILE INTEGRATED PEST MANAGEMENT (IPM)
Identifying pests of agriculture have relied heavily on experts, either extension researchers or agronomists trained in entomology, plant science, or plant pathology. Interactive keys are relatively new tools that allow for non-expert users to more easily identify species that are well known. Our team recently released a mobile app to help promote integrated pest management for Canadian field crops (www.mobile-ipm.com). The app functions to allow non-scientific users to identify crop pests, including insects and weeds. It also provides real-time forecasting of pest abundance levels and tools for managing crops that promote sustainable agriculture, including information on pesticide rotation and biological control. This product of this 5-year multi-collaborator project, and can be downloaded for free from Google Play and iTunes, will result in highly novel tool for sustainable agriculture.
CITIZEN SCIENCE AND POLLINATOR CONSERVATION:

The development of this app led to a new project funded from the Foundation for Food and Agriculture (FFAR) Pollinator Health Fund. This project builds upon the core software of Mobile-IPM to develop a nation-wide citizen science program to conserve our nation’s imperiled pollinators. Although in the early stages of development, my team is in the process of designing innovative games for average citizens to identify pollinators found in their yards and gardens. This includes using games combined with interactive identification technology and augmented reality. The software will be integrated with social media to attempt to develop a viral movement to convert lawns to pollinator habitat. More information can be found here: https://goo.gl/4FYbLi.

TAXONOMY & BIODIVERSITY
A more traditional aspect of my research involves describing life’s biota. While alpha taxonomy is often underappreciated, rarely cited, and less fashionable than evolutionary analyses, we cannot understand any aspect of biodiversity science if we do not know which species exist in different communities. I strive for open access to biodiversity data, and thus develop online digital keys and web-based information on local fauna with students. My passion for studying biodiversity naturally feeds the expansion and digitization of taxonomic collections, including the Central Florida Collection of Arthropods (Bug Closet) and the Wallis-Roughley Museum of Entomology (U of M).

BIOCONTROL AND SPECIES DELIMITATION
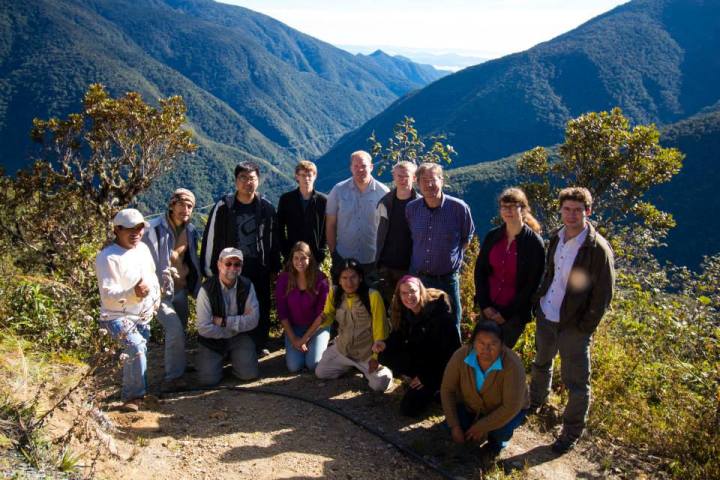
We are working with Dr. Toni Withers (Scion, NZL) and Dr. Geoff Allen (University of Tasmania) to examine a cryptic species complex of Eadya wasps (Braconidae). We discovered several new species (using integrative taxonomic approaches), one of which named Eadya daenerys is being investigated for biocontrol of the Eucalyptus Tortoise Beetle, Paropsis charybdis (Chrysomelidae).
Eadya daenerys sp.n. Ridenbaugh 2018
